DiscreteAliasUrn#
- class scipy.stats.sampling.DiscreteAliasUrn(dist, *, domain=None, urn_factor=1, random_state=None)#
Discrete Alias-Urn Method.
This method is used to sample from univariate discrete distributions with a finite domain. It uses the probability vector of size \(N\) or a probability mass function with a finite support to generate random numbers from the distribution.
- Parameters:
- distarray_like or object, optional
Probability vector (PV) of the distribution. If PV isn’t available, an instance of a class with a
pmfmethod is expected. The signature of the PMF is expected to be:def pmf(self, k: int) -> float. i.e. it should accept a Python integer and return a Python float.- domainint, optional
Support of the PMF. If a probability vector (
pv) is not available, a finite domain must be given. i.e. the PMF must have a finite support. Default isNone. WhenNone:If a
supportmethod is provided by the distribution object dist, it is used to set the domain of the distribution.Otherwise, the support is assumed to be
(0, len(pv)). When this parameter is passed in combination with a probability vector,domain[0]is used to relocate the distribution from(0, len(pv))to(domain[0], domain[0]+len(pv))anddomain[1]is ignored. See Notes and tutorial for a more detailed explanation.
- urn_factorfloat, optional
Size of the urn table relative to the size of the probability vector. It must not be less than 1. Larger tables result in faster generation times but require a more expensive setup. Default is 1.
- random_state{None, int,
numpy.random.Generator, numpy.random.RandomState}, optionalA NumPy random number generator or seed for the underlying NumPy random number generator used to generate the stream of uniform random numbers. If random_state is None (or np.random), the
numpy.random.RandomStatesingleton is used. If random_state is an int, a newRandomStateinstance is used, seeded with random_state. If random_state is already aGeneratororRandomStateinstance then that instance is used.
Methods
rvs([size, random_state])Sample from the distribution.
set_random_state([random_state])Set the underlying uniform random number generator.
Notes
This method works when either a finite probability vector is available or the PMF of the distribution is available. In case a PMF is only available, the finite support (domain) of the PMF must also be given. It is recommended to first obtain the probability vector by evaluating the PMF at each point in the support and then using it instead.
If a probability vector is given, it must be a 1-dimensional array of non-negative floats without any
infornanvalues. Also, there must be at least one non-zero entry otherwise an exception is raised.By default, the probability vector is indexed starting at 0. However, this can be changed by passing a
domainparameter. Whendomainis given in combination with the PV, it has the effect of relocating the distribution from(0, len(pv))to(domain[0],domain[0] + len(pv)).domain[1]is ignored in this case.The parameter
urn_factorcan be increased for faster generation at the cost of increased setup time. This method uses a table for random variate generation.urn_factorcontrols the size of this table relative to the size of the probability vector (or width of the support, in case a PV is not available). As this table is computed during setup time, increasing this parameter linearly increases the time required to setup. It is recommended to keep this parameter under 2.References
[1]UNU.RAN reference manual, Section 5.8.2, “DAU - (Discrete) Alias-Urn method”, http://statmath.wu.ac.at/software/unuran/doc/unuran.html#DAU
[2]A.J. Walker (1977). An efficient method for generating discrete random variables with general distributions, ACM Trans. Math. Software 3, pp. 253-256.
Examples
>>> from scipy.stats.sampling import DiscreteAliasUrn >>> import numpy as np
To create a random number generator using a probability vector, use:
>>> pv = [0.1, 0.3, 0.6] >>> urng = np.random.default_rng() >>> rng = DiscreteAliasUrn(pv, random_state=urng)
The RNG has been setup. Now, we can now use the
rvsmethod to generate samples from the distribution:>>> rvs = rng.rvs(size=1000)
To verify that the random variates follow the given distribution, we can use the chi-squared test (as a measure of goodness-of-fit):
>>> from scipy.stats import chisquare >>> _, freqs = np.unique(rvs, return_counts=True) >>> freqs = freqs / np.sum(freqs) >>> freqs array([0.092, 0.292, 0.616]) >>> chisquare(freqs, pv).pvalue 0.9993602047563164
As the p-value is very high, we fail to reject the null hypothesis that the observed frequencies are the same as the expected frequencies. Hence, we can safely assume that the variates have been generated from the given distribution. Note that this just gives the correctness of the algorithm and not the quality of the samples.
If a PV is not available, an instance of a class with a PMF method and a finite domain can also be passed.
>>> urng = np.random.default_rng() >>> class Binomial: ... def __init__(self, n, p): ... self.n = n ... self.p = p ... def pmf(self, x): ... # note that the pmf doesn't need to be normalized. ... return self.p**x * (1-self.p)**(self.n-x) ... def support(self): ... return (0, self.n) ... >>> n, p = 10, 0.2 >>> dist = Binomial(n, p) >>> rng = DiscreteAliasUrn(dist, random_state=urng)
Now, we can sample from the distribution using the
rvsmethod and also measure the goodness-of-fit of the samples:>>> rvs = rng.rvs(1000) >>> _, freqs = np.unique(rvs, return_counts=True) >>> freqs = freqs / np.sum(freqs) >>> obs_freqs = np.zeros(11) # some frequencies may be zero. >>> obs_freqs[:freqs.size] = freqs >>> pv = [dist.pmf(i) for i in range(0, 11)] >>> pv = np.asarray(pv) / np.sum(pv) >>> chisquare(obs_freqs, pv).pvalue 0.9999999999999999
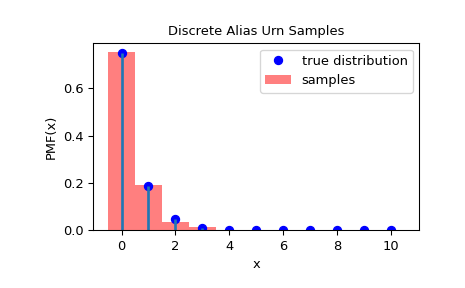
To check that the samples have been drawn from the correct distribution, we can visualize the histogram of the samples:
>>> import matplotlib.pyplot as plt >>> rvs = rng.rvs(1000) >>> fig = plt.figure() >>> ax = fig.add_subplot(111) >>> x = np.arange(0, n+1) >>> fx = dist.pmf(x) >>> fx = fx / fx.sum() >>> ax.plot(x, fx, 'bo', label='true distribution') >>> ax.vlines(x, 0, fx, lw=2) >>> ax.hist(rvs, bins=np.r_[x, n+1]-0.5, density=True, alpha=0.5, ... color='r', label='samples') >>> ax.set_xlabel('x') >>> ax.set_ylabel('PMF(x)') >>> ax.set_title('Discrete Alias Urn Samples') >>> plt.legend() >>> plt.show()
To set the
urn_factor, use:>>> rng = DiscreteAliasUrn(pv, urn_factor=2, random_state=urng)
This uses a table twice the size of the probability vector to generate random variates from the distribution.