estimated_cdf#
- scipy.stats.estimated_cdf(x, y, *, method='linear', axis=0, nan_policy='propagate', keepdims=None)[source]#
Estimate the CDF of the distribution underlying a sample.
- Parameters:
- xarray_like of real numbers
Data array.
- yarray_like of real numbers
Points at which to evaluate the estimated cdf. See axis, keepdims, and the examples for broadcasting behavior.
- methodstr, default: ‘linear’
The method to use for estimating the cumulative distribution function. The available options, numbered as they appear in [1], are:
‘inverted_cdf’ (AKA the empirical cumulative distribution function)
‘averaged_inverted_cdf’
‘closest_observation’
‘interpolated_inverted_cdf’
‘hazen’
‘weibull’
‘linear’ (default)
‘median_unbiased’
‘normal_unbiased’
See Notes for details.
- axisint or None, default: 0
Axis along which samples in x are given in ND case.
Noneravels both x and y before performing the calculation, without checking whether the original shapes were compatible. As in otherscipy.statsfunctions, a positive integer axis is resolved after prepending 1s to the shape of x or y as needed until the two arrays have the same dimensionality. When providing x and y with different dimensionality, consider using negative axis integers for clarity.- nan_policystr, default: ‘propagate’
Defines how to handle NaNs in the input data x.
propagate: if a NaN is present in the axis slice (e.g. row) along which the statistic is computed, the corresponding slice of the output will contain NaN(s).omit: NaNs will be omitted when performing the calculation. If insufficient data remains in the axis slice along which the statistic is computed, the corresponding slice of the output will contain NaN(s).raise: if a NaN is present, aValueErrorwill be raised.
If NaNs are present in y, the corresponding entries in the output will be NaN.
- keepdimsbool, optional
Consider the case in which x is 1-D and y is a scalar: the estimated cumulative distribution function at a point is a reducing statistic, and the default behavior is to return a scalar. If keepdims is set to True, the axis will not be reduced away, and the result will be a 1-D array with one element.
The general case is more subtle, since multiple values of y may be specified for each axis-slice of x. For instance, if both x and y are 1-D and
y.size > 1, no axis can be reduced away; there must be an axis to contain the number of values given byy.size. Therefore:By default, the axis will be reduced away if possible (i.e. if there is exactly one element of y per axis-slice of x).
If keepdims is set to True, the axis will not be reduced away.
If keepdims is set to False, the axis will be reduced away if possible, and an error will be raised otherwise.
- Returns:
- probabilityscalar or ndarray
The resulting probabilities(s). The dtype is the result dtype of x, y, and the Python
floattype.
See also
Notes
Given a sample x from an underlying distribution,
estimated_cdfprovides a nonparametric estimate of the cumulative distribution function.By default, this is done by computing the “fractional index”
pat whichywould appear withinz, a sorted copy of x:p = 1 / (n - 1) * (j + ( y - z[j]) / (z[j+1] - z[j]))
where the index
jis that of the largest element ofzthat does not exceedy, andnis the number of elements in the sample. Note that ifyis an element ofz, thenjis the index such thaty = z[j], and the formula simplifies to the intuitivej / (n - 1). The full formula linearly interpolates betweenj / (n - 1)and(j + 1) / (n - 1).This is a special case of the more general:
p = (j + (y - z[j]) / (z[j+1] - z[j] + 1 - m) / n
where
mmay be defined according to several different conventions. The preferred convention may be selected using themethodparameter:methodnumber in H&F
minterpolated_inverted_cdf4
0hazen5
1/2weibull6
plinear(default)7
1 - pmedian_unbiased8
p/3 + 1/3normal_unbiased9
p/4 + 3/8Note that indices
jandj + 1are clipped to the range0ton - 1when the results of the formula would be outside the allowed range of non-negative indices, and resultingpis clipped to the range0to1.The table above includes only the estimators from [1] that are continuous functions of probability p (estimators 4-9). SciPy also provides the three discontinuous estimators from [1] (estimators 1-3), where
jis defined as above,mis defined as follows, andgis0whenindex = p*n + m - 1is less than0and otherwise is defined below.inverted_cdf:m = 0andg = int(index - j > 0)averaged_inverted_cdf:m = 0andg = (1 + int(index - j > 0)) / 2closest_observation:m = -1/2andg = 1 - int((index == j) & (j%2 == 1))
When all the data in
xare unique,estimated_cdfandquantileare are inverses of one another within a certain domain. Althoughquantilewithmethod='linear'is invertible over the whole domain ofpfrom0to1, this is not true of other methods.Array API Standard Support
estimated_cdfhas experimental support for Python Array API Standard compatible backends in addition to NumPy. Please consider testing these features by setting an environment variableSCIPY_ARRAY_API=1and providing CuPy, PyTorch, JAX, or Dask arrays as array arguments. The following combinations of backend and device (or other capability) are supported.Library
CPU
GPU
NumPy
✅
n/a
CuPy
n/a
✅
PyTorch
✅
✅
JAX
✅
✅
Dask
⛔
n/a
estimated_cdfalso accepts MArrays backed by the backends indicated above; masked values will be treated as though they were not present.See Support for the array API standard for more information.
References
Examples
>>> import numpy as np >>> from scipy import stats >>> x = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
Estimate the cumulative distribution function of one sample at a single point.
>>> stats.estimated_cdf(x, 5, axis=-1) np.float64(0.5)
Estimate the cumulative distribution function of one sample at two points.
>>> stats.estimated_cdf(x, [2.5, 7.5], axis=-1) array([0.25, 0.75])
Estimate the cumulative distribution function of two samples at different points.
>>> x = np.stack((np.arange(0, 11), np.arange(10, 21))) >>> stats.estimated_cdf(x, [[2.5], [17.5]], axis=-1, keepdims=True) array([[0.25], [0.75]])
Estimate the cumulative distribution function at many points for each of two samples.
>>> import matplotlib.pyplot as plt >>> rng = np.random.default_rng() >>> x = stats.Normal(mu=[-1, 1]).sample(10000) >>> y = np.linspace(-4, 4, 5000)[:, np.newaxis] >>> p = stats.estimated_cdf(x, y, axis=0) >>> plt.plot(y, p) >>> plt.show()
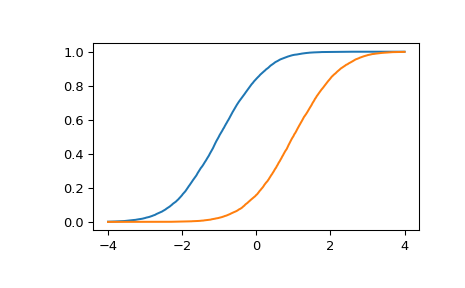
Note that the
quantileandestimated_cdffunctions are inverses of one another within a certain domain.>>> p = np.linspace(0, 1, 300) >>> x = rng.standard_normal(300) >>> y = stats.quantile(x, p) >>> p2 = stats.estimated_cdf(x, y) >>> np.testing.assert_allclose(p2, p) >>> y2 = stats.quantile(x, p2) >>> np.testing.assert_allclose(y2, y)
However, the domain over which
quantilecan be inverted byestimated_cdfdepends on the method used. This is most noticeable when there are few observations in the sample.>>> x = np.asarray([0, 1]) >>> y_linear = stats.quantile(x, p, method='linear') >>> y_weibull = stats.quantile(x, p, method='weibull') >>> y_iicdf = stats.quantile(x, p, method='interpolated_inverted_cdf') >>> plt.plot(p, y_linear, p, y_weibull, p, y_iicdf) >>> plt.legend(['linear', 'weibull', 'iicdf']) >>> plt.xlabel('p') >>> plt.ylabel('y = quantile(x, p)') >>> plt.show()
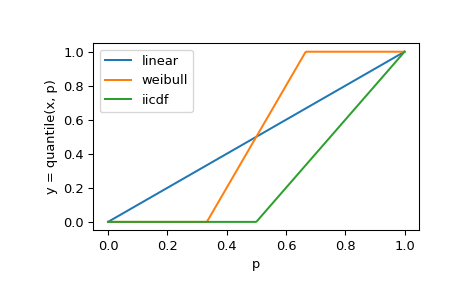
For example, in the case above,
quantileis only invertible fromp = 0.5top = 1.0withmethod = 'interpolated_inverted_cdf'. This is a fundamental characteristic of the methods, not a shortcoming ofestimated_cdf.