ev#
- LSQSphereBivariateSpline.ev(theta, phi, dtheta=0, dphi=0)[source]#
Evaluate the spline at points
Returns the interpolated value at
(theta[i], phi[i]), i=0,...,len(theta)-1.- Parameters:
- theta, phiarray_like
Input coordinates. Standard Numpy broadcasting is obeyed. The ordering of axes is consistent with np.meshgrid(…, indexing=”ij”) and inconsistent with the default ordering np.meshgrid(…, indexing=”xy”).
- dthetaint, optional
Order of theta-derivative
Added in version 0.14.0.
- dphiint, optional
Order of phi-derivative
Added in version 0.14.0.
Examples
Suppose that we want to use splines to interpolate a bivariate function on a sphere. The value of the function is known on a grid of longitudes and colatitudes.
>>> import numpy as np >>> from scipy.interpolate import RectSphereBivariateSpline >>> def f(theta, phi): ... return np.sin(theta) * np.cos(phi)
We evaluate the function on the grid. Note that the default indexing=”xy” of meshgrid would result in an unexpected (transposed) result after interpolation.
>>> thetaarr = np.linspace(0, np.pi, 22)[1:-1] >>> phiarr = np.linspace(0, 2 * np.pi, 21)[:-1] >>> thetagrid, phigrid = np.meshgrid(thetaarr, phiarr, indexing="ij") >>> zdata = f(thetagrid, phigrid)
We next set up the interpolator and use it to evaluate the function at points not on the original grid.
>>> rsbs = RectSphereBivariateSpline(thetaarr, phiarr, zdata) >>> thetainterp = np.linspace(thetaarr[0], thetaarr[-1], 200) >>> phiinterp = np.linspace(phiarr[0], phiarr[-1], 200) >>> zinterp = rsbs.ev(thetainterp, phiinterp)
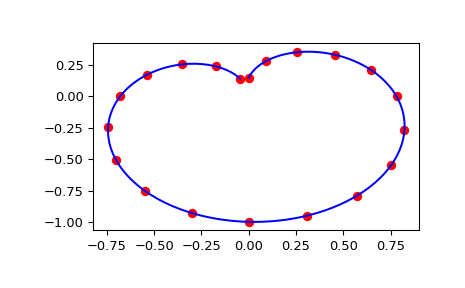
Finally we plot the original data for a diagonal slice through the initial grid, and the spline approximation along the same slice.
>>> import matplotlib.pyplot as plt >>> fig = plt.figure() >>> ax1 = fig.add_subplot(1, 1, 1) >>> ax1.plot(np.sin(thetaarr) * np.sin(phiarr), np.diag(zdata), "or") >>> ax1.plot(np.sin(thetainterp) * np.sin(phiinterp), zinterp, "-b") >>> plt.show()